HOME | satellite telemetry | acoustic telemetry | geolocators | radio telemetry | motus wildlife tracking system
individual marking | molecular markers | stable isotopes | movement models | future methods
Overview
Molecular genetic analyses are commonly used to study dispersal and population connectivity in animals. Innovative use of genetic data, particularly in combination with other approaches, has provided new insights on movement from local to global scales. Topics of interest include sex-biased dispersal patterns, natal site fidelity and dispersal trends, long-distance dispersal, movement or gene flow between populations, and identification of migration pathways (Haig et al. 2011).
In the past, investigations have not always yielded the most definitive results because it was difficult to identify enough markers capable of differentiating populations. However, recent development of fast-throughput sequencing has revolutionized our ability to identify hundreds of thousands of variable microsatellites and other types of genetic markers (see Lerner and Fleischer 2010).
What are they?
Molecular markers, such as DNA markers, in migratory connectivity research are used to identify genetic differentiation among populations. Markers include mitochondrial (mt)DNA and microsatellites that may be different throughout the range of a species (Broquet et al. 2009, Lerner and Fleischer 2010, Lowe and Allendorf 2010).
Tracking populations throughout the annual cycle
Birds and other flying animals that readily disperse hundreds or thousands of miles to new breeding areas often have panmictic populations that do not show any evidence of genetic differences (Broquet and Petit 2009). This can make it difficult to use molecular markers to track populations throughout the annual cycle. There are three potential solutions to this problem:
- Instead of randomly selecting variable low-quality markers from a small resource group to screen populations, we can selectively choose markers from a large pool to better optimize our resolution and ability to differentiate populations (as in Haig et al. 1997). This approach can provide genetic markers capable of high resolution at the population level, which can be used to monitor movements of different populations throughout the year.
- Combine molecular genetic analyses with results from stable isotopes and habitat suitability modeling. Molecular markers tend to be better at differentiating east-west patterns while isotopes are better at discerning north-south patterns. Together they can be a powerful tool in tracking animal movements and understanding migratory connectivity. See Ruegg et al. (2017) and Veen et al. (2013) for more details.
- Use a surrogate for the species of interest. Population-specific markers for a parasite or other organism that accompanies an animal in its movements can be molecularly sampled (Fallon et al. 2006). Bensch and Akesson (2003) demonstrated this approach by sampling parasites found on Willow Warblers from various populations. They illustrated that the parasite’s DNA yielded a more refined understanding of the host species movement patterns than the host’s DNA itself.
Although analyses of high-dispersal species may have a number of challenges, it is important to recognize that many widely dispersing species have populations that are readily identifiable with molecular markers. Most noteworthy are those where geneticists have used this attribute to track illegal possession of marine mammal meat (see Baker et al. 2008, 2010).
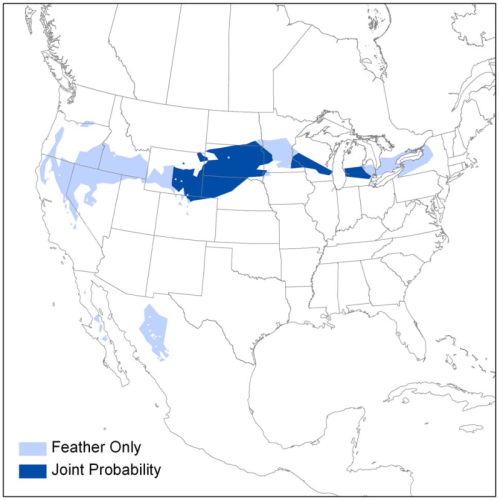
The combination of stable isotopes and genetic markers can improve location estimates. An example of the combination of stable hydrogen isotopes and genetic information reduces probable origin area of Loggerhead Shrikes in Chabot et al. (2012).
Collection

Museum collections are useful for understanding how genetics have have changed over time. Holmes et al (2016) show this relationship here.
Analysis of molecular markers in individuals requires the collection of genetic material. This is most commonly collected through blood samples, but can also include hair or feathers. mRNA extracts can also come from tissues such as brain, eye, or liver (Liedvogel et al. 2011).
A benefit of using this method is that samples can also be collected from museum specimens. Genetic materials collected from museum collections can help answer questions about the evolution of traits, including migration and migratory connectivity (Holmes et al. 2016). The use of ancient DNA in particular can reveal historical patterns and connectivity of species to better understand current patterns and predict future changes (Beadell et al. 2009, Hoelzel 2010).
Analysis
The strongest connectivity studies are carried out using a variety of approaches. Often a molecular component would be useful but a scientist is not set up to carry out the analyses in their own lab. Fortunately, many labs routinely carry out molecular analyses of population connectivity and are happy to discuss partnerships. Working with service labs requires forming a scientific partnership.
Edited by Susan Haig (USGS Forest and Rangeland Ecosystem Science Center, susan_haig@usgs.gov), 2014.
Updated by Allison Huysman (Migratory Connectivity Project, huysmana@si.edu), 2020.
References
- Baker, C.S., D.J. Steel, Y. Choi, H. Lee, K.S. Kim, S.K. Choi, Y. Ma, C. Hambleton, L. Psihoyos, R.L. Brownell et al. 2010. Genetic evidence of illegal trade in protected whales links Japan with the US and South Korea. Biology Letters.
- Baker, C.S. 2008. A truer measure of the market: the molecular ecology of fisheries and wildlife trade. Molecular Ecology 17:3985-3998.
- Beadell, J, Y. Chan and R. Fleischer. 2009. The role of ancient DNA in conservation biology. Pages 202-224 in: Population Genetics for Animal Conservation. (eds G. Bertorelle, M. W. Bruford, H. C. Hauffe, A. Rizzoli and C. Vernesi), Cambridge University Press.
- Bensch, S., and S. Akesson. 2003. Temporal and spatial variation in Hematozoans in Scandinavian Willow Warblers. Journal of Parasitology 88: 388-391.
- Broquet, T., and E. J. Petit. 2009. Molecular estimation of dispersal for ecology and population genetics. Annual Review of Ecology Evolution and Systematics 40:193-216.
- Broquet, T., J. Yearsley, A.H. Hirzel, J. Goudet, and N. Perrin. 2009. Inferring recent migration rates from individual genotypes. Molecular Ecology 18:1048-1060.
- Chabot, A.A., K.A. Hobson, S.L. Van Wilgenburg, G.J. McQuat, and S.C. Lougheed. 2012. Advances in linking wintering migrant birds to their breeding-ground origins using combined analyses of genetic and stable isotope markers. PLoS ONE 7(8): e43627.
- Fallon, S. M., R. C. Fleischer, and G. R. Graves. 2006. Malarial parasites as geographical markers in migratory birds? Biology Letters 2:213-216.
- Haig, S.M., W.M. Bronaugh, R.S. Crowhurst, J. D’Elia, C.A. Eagles-Smith, C.W. Epps, B. Knaus, M.P. Miller, M.L. Moses, S. Oyler-McCance, W.D. Robinson, and B. Sidlauskas. 2011. Genetic applications in avian conservation. The Auk 128(2): 205-229.
- Haig, S.M., C.L. Gratto-Trevor, T.D. Mullins, and M.A. Colwell. 1997. Population identification of western hemisphere shorebirds throughout the annual cycle. Molecular Ecology 6: 413-427.
- Hoelzel, A. R. 2010. Looking backward to look forwards: conservation genetics in a changing world. Conservation Genetics 11:655-660.
- Holmes, M.W., T.T. Hammond, G.O.U. Wogan, R.E. Walsh, K. Labarbera, E.A. Wommack, F.M. Martins, J.C. Crawford, K.L. Mack, L.M. Bloch, and M.W. Nachman. 2016. Natural history collections as windows on evolutionary processes. Molecular Ecology 25:864-881.
- Lerner, H., and R. Fleischer. 2010. Prospects for the use of next-generation sequencing methods in ornithology. Auk 127:4-15.
- Liedvogel, M., S. Åkesson, and S. Bensch. 2011. The genetics of migration on the move. Trends in Ecology and Evolution 26: 561-569.
- Lowe, W. H., and F. W. Allendorf. 2010. What can genetics tell us about population connectivity? Molecular Ecology 19:3038-3051.
- Ruegg, K.C., E.C. Anderson, R.J. Harrigan, K.L. Paxton, J.F. Kelly, F. Moore, and T.B. Smith. 2017. Genetic assignment with isotopes and habitat suitability (GAIAH), a migratory bird case study. Methods in Ecology and Evolution 8: 1241-1252.
- Veen, T. 2013. Unravelling migratory connections: the next level. Molecular Ecology 22: 4144-4146.
HOME | satellite telemetry | acoustic telemetry | geolocators | radio telemetry | motus wildlife tracking system
individual marking | molecular markers | stable isotopes | movement models | future methods


